User Tools
Sidebar
Table of Contents
Selection Manager
1. Introduction
VMD is an exceptional program to visualize molecular structures. The main drawback of this software is the learning curve that turns it hard to use by non-expert users. One of the major difficulties that non-expert users have while handling VMD regards the selection of atoms. This is done through a series of keywords that the user must dominate in order to for example visualize and/or hide particular parts of the molecule, do specific transformations on the atoms of that selectin, etc.
In order to overcome this issues we have developed “Selection Manager”. This plug-in allows to do common selections on molecules without requiring any prior knowledge regarding the keywords available on VMD. It makes the selection of atoms simpler and easy for non-expert users. In addition it still allows the insertion of advanced commands but alerts the user for any error that might exist on the syntax.
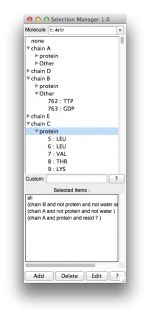
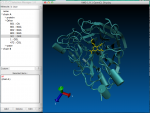
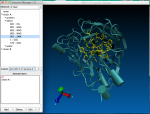
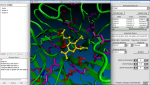
2. ScreenShots
3. Download
In order to run the”Selection Manager” plug-in, the Visual Molecular dynamics software (VMD) must be installed. It can be found at http://www.ks.uiuc.edu/Research/vmd/ .
4. Installation
The selection manager plug-in can be installed in any operating systems where VMD works. In order to work the user has to add the following lines to the .vmdrc file that can be found in the home directory:
#### selectionMAnager lappend auto_path /home/users/nunocerqueira/selectionManager vmd_install_extension selV "::selV::start" "PortoBioComp/Selection Manager" ## END
Note: /home/users/nunocerqueira/selectionManager is the directory where the plug-in was installed.
5. Tutorial
not available yet.