Manganese(II) - DEE[SO2]
Crystal structures from Mandelate Racemase show that it is octameric.
The manganese(II) atom in this enzyme is bonded to the side chains of an aspartate, two glutamates and it is bidentate either to a sulfate ion or a S-altrolactate.
| COORDINATION SPHERE |
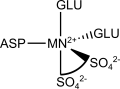 |
Oxidation State: Mn(II)
Spin Multiplicity: 6 |
Structure chosen to parameterize
| TEST PROTEIN |
| Protein | Mandelate Racemase |
| PDB Code | 2MNR |
| Crystallographic Resolution | 1.90 Å |
| Organism | Pseudomonas putida |
| [Neidhart, 1991] |
Parameters Determined
Atom Types
| ATOM TYPE | NEW ATOM TYPE | MASS | STRUCTURE |
| MN | M1 | 54.938 | 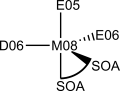 |
| OD2 | D1 | 15.9994 |
| OE2 | E1 | 15.9994 |
| OE1 | E2 | 15.9994 |
Bond Parameters
| BOND | Kl / kcal mol-1 Å-2 | l / Å | REFERENCE |
| M1-D1 | 33.32 | 2.2509 | [Neves, 2013] |
| M1-E1 | 47.15 | 2.1737 |
| M1-E2 | 49.51 | 2.2008 |
| M1-R1 | 41.81 | 2.1619 |
| M1-R2 | 45.80 | 2.1526 |
| D1-C | 656.0 | 1.25 | |
| E1-C | 656.0 | 1.25 | |
| E2-C | 656.0 | 1.25 | |
| SO-R1 | 541.1 | 1.436 | |
| SO-R2 | 541.1 | 1.436 | |
| SO-O1 | 541.1 | 1.436 | |
| SO-O2 | 541.1 | 1.436 | |
Angle Parameters
| ANGLE | Kθ / kcal mol-1 rad-2 | θ / deg | REFERENCE |
| D1-M1-E1 | 45.46 | 92.3315 | [Neves, 2013] |
| D1-M1-E2 | 25.03 | 123.0771 |
| D1-M1-R1 | 13.27 | 96.5510 |
| D1-M1-R2 | 4.321 | 121.0757 |
| E1-M1-E2 | 40.08 | 86.7147 |
| E1-M1-R1 | 19.90 | 160.9449 |
| E1-M1-R2 | 33.00 | 93.3926 |
| E2-M1-R1 | 8.95 | 102.2828 |
| E2-M1-R2 | 4.405 | 115.7857 |
| R1-M1-R2 | 291.6 | 67.5889 |
| M1-D1-C | 16.912 | 165.3084 |
| M1-E1-C | 22.912 | 145.6796 |
| M1-E2-C | 15.527 | 135.3597 |
| M1-R1-SO | 100.89 | 95.8499 |
| M1-R2-SO | 100.89 | 95.8499 |
| D1-C-CT | 70.0 | 117.0 | |
| D1-C-O2 | 80.0 | 126.0 | |
| E1-C-CT | 70.0 | 117.0 | |
| E1-C-O2 | 80.0 | 126.0 | |
| E2-C-CT | 70.0 | 117.0 | |
| E2-C-O2 | 80.0 | 126.0 | |
| R1-SO-O1 | 74.6 | 119.82 | |
| R1-SO-O2 | 74.6 | 119.82 | |
| R1-SO-R2 | 74.6 | 119.82 | |
| R2-SO-O1 | 74.6 | 119.82 | |
| R2-SO-O2 | 74.6 | 119.82 | |
| O1-SO-O2 | 74.6 | 119.82 | |
Van der Waals Parameters
| ATOM TYPE | Ri / Å | εi / kcal mol-1 | REFERENCE |
| M1 | 1.4544 | 0.03 | [Babu, 2006] |
| D1 | 1.6612 | 0.21 | |
| E1 | 1.6612 | 0.21 | |
| E2 | 1.6612 | 0.21 | |
| R1 | 1.6612 | 0.2100 | |
| R2 | 1.6612 | 0.2100 | |
| O1 | 1.6612 | 0.2100 | |
| O2 | 1.6612 | 0.2100 | |
| SO | 2.0000 | 0.2500 | |
Validation of Parameters from MD Simulations
Bond Parameters
| BOND | l0 crystal | l0 opt | ‹l›MD ± σ(l) | ‹l›MD ± 2σ(l) |
| M1-D1 | 2.03 | 2.25 | 2.41 ± 0.09 | [2.318; 2.506] |
| M1-E1 | 2.15 | 2.17 | 2.28 ± 0.08 | [2.200; 2.359] |
| M1-E2 | 1.95 | 2.20 | 2.27 ± 0.08 | [2.193; 2.347] |
| M1-R2 | 3.54 | 2.16 | 2.18 ± 0.06 | [2.114; 2.238] |
| M1-R2 | 2.20 | 2.15 | 2.17 ± 0.06 | [2.113; 2.232] |
Angle Parameters
| ANGLE | θ0 crystal | θ0 QM | ‹θ›MD ± σ(θ) | ‹θ›MD ± 2σ(θ) |
| D1-M1-E1 | 87.7 | 92.3 | 89 ± 4 | [80.5; 97.0] |
| D1-M1-E2 | 170.0 | 123.1 | 127 ± 5 | [117.4; 137.5] |
| D1-M1-R1 | 85.6 | 96.6 | 91 ± 6 | [78.7; 103.7] |
| D1-M1-R2 | 94.6 | 121.1 | 99 ± 8 | [82.2; 115.1] |
| E1-M1-E2 | 88.0 | 86.7 | 87 ± 5 | [78.0; 96.9] |
| E1-M1-R1 | 129.4 | 160.9 | 156 ± 4 | [147.4; 163.8] |
| E1-M1-R2 | 92.1 | 93.4 | 92 ± 4 | [83.3; 100.7] |
| E2-M1-R1 | 104.1 | 102.3 | 111 ± 6 | [97.8; 123.6] |
| E2-M1-R2 | 94.6 | 115.8 | 133 ± 9 | [115.9; 150.5] |
| R1-M1-R2 | 39.0 | 67.6 | 66 ± 1 | [63.1; 69.0] |
Downloads
| Parameter File | .frcmod |
| Coordination Sphere Charges | .off |
Mandelate Racemase’s PDB File (2MNR.pdb)
LEAPRC file | .pdb, leaprc |
References

